Differential Scanning Fluorimetry Reaches New Peaks on the NISTmAb
Introduction
The NISTmAb is a widely characterised monoclonal antibody intended to be used as a reference molecule in the development of novel technology for therapeutic protein characterisation. In this application note, we use the SUPR-DSF to acquire differential scanning fluorimetry (DSF) data on the NISTmAb and compare the results against differential scanning calorimetry (DSC).
The SUPR-DSF resolved all three domains of the NISTmAb: CH2 (69°C), CH3 (83°C), and the unusually high melting temperature of the Fab domain (94°C).[ 1] In addition, the apparent melting temperatures agreed exceptionally well with the literature results. These results validate you can easily obtain high quality protein stability information with the SUPR-DSF.
Results
The fluorescence spectra from the SUPR-DSF thermal melt experiment were used to calculate the barycentric mean (BCM) to quantify the wavelength shifts of the NISTmAb as it transitioned from the folded to the unfolded state. The NISTmAb BCM melt curve shown in Figure 1(a) illustrates three distinct transitions: CH2 (69°C), CH3 (83°C), and Fab (94°C) domains. The SUPR-DSF can accurately measure the unusually high Fab melting temperature due to its ability to ramp samples up to 105°C, which is not easily possible by alternative DSF platforms. This is evident in Figure 1(a) where the alternative DSF platform is unable to completely resolve the Fab transition.[2]
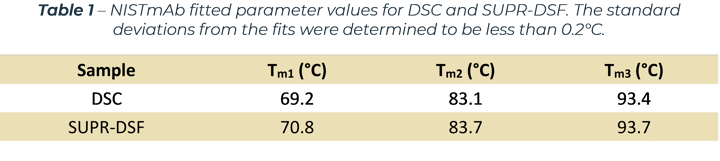
The SUPR-DSF BCM curve was processed with a first derivative and compared to the DSC data by normalising to the Fab domain.[1] The result, shown in Figure 2(b), clearly demonstrates excellent agreement between the two measurements depicted by all three peaks aligning with each other. The excellent correlation between the techniques shows the intrinsic protein fluorescence provides an orthogonal approach for measuring protein stability. This opens new possibilities by being able to obtain high quality melting curves on 384 samples in under 2 hours with the SUPR‑DSF. The melting temperatures calculated by both methods are shown in Table 1.
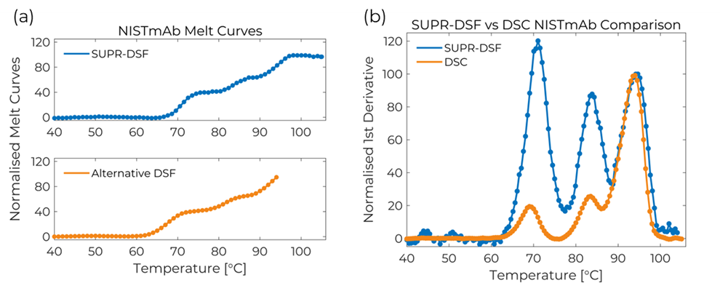
Figure 1 – (a) SUPR-DSF BCM melt curve for NISTmAb compared to alternative DSF platform. (b) Normalised first derivative of BCM data compared against DSC results.

Conclusion
The SUPR-DSF from Protein Stable can quickly and accurately measure melting temperatures by monitoring the wavelength shifts of intrinsic protein fluorescence. Results shown here have demonstrated the SUPR-DSF can obtain high quality, accurate melting temperatures of the NISTmAb, including the unusually high melting temperature of the Fab domain (94°C). This validates the intrinsic fluorescence based DSF method of the SUPR-DSF can achieve the same results as DSC while increasing throughput, convenience, and lower sample consumption by taking advantage of 384-well microplates.
Methodology
NISTmAb was prepared at 1 mg⁄mL in L-Histidine pH 6.0 buffer and dispensed in a 384‑well PCR microplate at a well volume of 10 µL. After dispensing, the plate was centrifuged for 60 seconds and sealed with an optically clear adhesive film. The SUPR-DSF was set up to measure the fluorescence spectra from 15°C to 105°C with a 1 °C per minute ramp rate.
Fluorescence spectra were processed with BCM calculation to quantify the spectral shift. The BCM data was analysed as a function of temperature and processed with a first derivative. Melting temperatures were calculated from the peaks of the derivative. Methods and results for all samples are available on request.
References
[ 1] – Y. Gokarn, S. Agarwal, K Arthur, A Bepperling, et al. “Biophysical Techniques for Characterizing the Higher Order Structure and Interactions of Monoclonal Antibodies,” in State-of-the-Art and Emerging Technologies for Therapeutic Monoclonal Antibody Characterization Volume 2. Biopharmaceutical Characterization: The NISTmAb Case Study. Chapter 6, p.285-327.2015
[2] – Y. Hamuro, M. Derebe, S. Venkataramani, J. Nemeth. “The Effects of Intramolecular and Intermolecular Electrostatic Repulsions on the Stability and Aggregation of NISTmAb Revealed by HDX-MS, DSC and NanoDSF”. Protein Science. Vol. 30, Issue 8, p. 1686-1700. 2021.
Download PDF
